Publications
Papers submitted to refereed journals
Papers in refereed journals
[1] Kevin Sutanto and Marcel Turcotte. Assessing global-local secondary structure fingerprints to classify
RNA sequences with deep learning. IEEE/ACM Transactions on Computational Biology and Bioinformatics
Special Issue, Submitted on 2021-02-28; Major Revision Requested on 2021-05-12; Accepted for publication on
2021-09-29.
[2] Aseel Awdeh, Marcel Turcotte, and Theodore J. Perkins. WACS: improving ChIP-seq peak calling by
optimally weighting controls. BMC Bioinformatics, 22(1):69, 2021.
[3] Alexander Gawronski and Marcel Turcotte. RiboFSM: Frequent subgraph mining for the discovery of
RNA structures and interactions. BMC Bioinformatics, 15 Suppl 13:S2, November 2014.
[4] Ghada Badr, Isra Al-Turaiki, Marcel Turcotte, and Hassan Mathkour. IncMD: Incremental trie-based
structural motif discovery algorithm. Journal of Bioinformatics and Computational Biology, 12(05):1450027,
October 2014. PMID: 25362841.
[5] Georgette Kiethega, Yifei Yan, Marcel Turcotte, and Gertraud Burger. RNA-level unscrambling of
fragmented genes in Diplonema mitochondria. RNA biology, 10(2):301–313, January 2013.
[6] Georgette N Kiethega, Marcel Turcotte, and Gertraud Burger. Evolutionarily Conserved cox1
Trans-Splicing Without cis-Motifs. Molecular biology and evolution, 28(9):2425–2428, September 2011.
[7] Mikhail Jiline, Stan Matwin, and Marcel Turcotte. Annotation Concept Synthesis and Enrichment
Analysis: a Logic-Based Approach to the Interpretation of High-Throughput Experiments. Bioinformatics,
27(17):2391–2398, July 2011.
[8] A. Bellamy-Royds and M. Turcotte. Can clustal-style progressive pairwise alignment of multiple
sequences be applied to RNA secondary structure prediction? BMC Bioinformatics, 8:190, 2007.
[9] V. X. Jin and M. Turcotte. Detecting localized interspersed motifs in genomic sequences: Application to
the mouse genome. IEEE Transactions on Instrumentation and Measurement, 56(5):1770–1775, October 2007.
[10] S. D. Baird, S. M. Lewis, M. Turcotte, and M. Holcik. A search for structurally similar cellular internal
ribosome entry sites. Nucl. Acids Res., 35(14):4664–4677, 2007.
[11] S. D. Baird, M. Turcotte, R. G. Korneluk, and M. Holcik. Searching for IRES. RNA Journal,
12(10):1755–1785, 2006.
[12] M. Anwar, T. Nguyen, and M. Turcotte. Identification of consensus RNA secondary structures using
suffix arrays. BMC Bioinformatics, 7:244, 2006.
[13] B. Masoumi and M. Turcotte. Simultaneous alignment and structure prediction of three RNA sequences.
International Journal of Bioinformatics Research and Applications, 1(2):230–245, 2005.
[14] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. Generating protein three-dimensional fold signatures
using inductive logic programming. Computers & Chemistry, 26(1):57–64, December 2001.
[15] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. Automated discovery of structural signatures of
protein fold and function. J. Mol. Biol., 306(3):591–605, 2001.
[16] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. The effect of relational background knowledge on
learning of protein three-dimensional fold signatures. Machine Learning, 43:81 – 95, 2001.
[17] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. Use of inductive logic programming to learn principles
of protein structure. Electronic Transactions on Artificial Intelligence, ETAI, 5(39), 2000.
[18] S. A. Benner, G. Cannarozzi, D. Gerloff, M. Turcotte, and G. Chelvanayagam. Bona Fide prediction
of protein secondary structure using transparent analyses of multiple sequence alignments. Chemical Review,
97(8):2725 – 2843, 1997.
[19] D.L. Gerloff, F.E. Cohen, C. Korostensky, M. Turcotte, G.H. Gonnet, and S.A. Benner. A predicted
concensus for the N-terminal fragment of the heat shock protein HSP90 family. Proteins: Struct. Funct. Genet.,
27:450–458, 1996.
[20] M. Turcotte, G. Lapalme, and F. Major. Exploring the conformations of nucleic acids. J. Funct. Prog.,
5(3):443–460, 1995.
[21] M. Feeley, M. Turcotte, and G. Lapalme. Using Multilisp for solving constraint satisfaction problems:
an application to nucleic acid 3D structure determination. LISP AND SYMBOLIC COMPUTATION,
7(2/3):232–247, 1994.
[22] F. Major, M. Turcotte, , D. Gautheret, G. Lapalme, E. Fillion, and R. Cedergren. The combination
of symbolic and numerical computation for three-dimensional modeling of RNA. Science, 253:1255–1260,
September 1991.
Papers in refereed conferences submitted
Papers in referred conference proceedings
[1] Kevin Sutanto and Marcel Turcotte. Extracting and evaluating features from RNA virus sequences to
predict host species susceptibility using deep learning. In 13th International Conference on Bioinformatics and
Biomedical Technology (ICBBT 2021), Northwestern Polytechnical University, Xi’an, China, May 21-23 2021.
[2] Kevin Sutanto and Marcel Turcotte. Assessing the use of secondary structure fingerprints and deep
learning to classify RNA sequences. In IEEE International Conference on Bioinformatics and Biomedicine
(BIBM), Seoul, South Korea, December 16-19 2020.
[3] Manuel Belmadani and Marcel Turcotte. MotifGP: Using multi-objective evolutionary computing for
mining network expressions in DNA sequences. In IEEE International Conference on Computational Intelligence
in Bioinformatics and Computational Biology (CIBCB 2016), Chiang Mai, Thailand, October, 5-7, 2016 2016.
[4] Isra Al-Turaiki, Ghada Badr, Marcel Turcotte, and Hassan Mathkour. Incremental structural motif
discovery. In International Symposium on Bioinformatics Research and Applications (ISBRA), 2013.
[5] Alexander Gawronski and Marcel Turcotte. Novel framework for the discovery of RNA elements and its
application to Euglenozoa. In International Symposium on Bioinformatics Research and Applications (ISBRA),
2013.
[6] Oksana Korol and Marcel Turcotte. Learning relationships between over-represented motifs in a set
of DNA sequences. In CIBCB 2012 : IEEE Symposium on Computational Intelligence in Bioinformatics and
Computational Biology, San Diego, USA, May 9–12 2012.
[7] Sandrine Moreira, Marcel Turcotte, and Gertraud Burger. NGS: Neglected genome sequencing assembly
and annotation challenges in a highly divergent protozoan genome. In JOBIM 2011 : Journées Ouvertes en
Biologie, Informatique et Mathématiques, Paris, France, June 28 – July 1 2011.
[8] Ghada Badr and Marcel Turcotte. Component-based matching for multiple interacting rna sequences.
In Jianer Chen, Jianxin Wang, and Alexander Zelikovsky, editors, Bioinformatics Research and Applications,
volume 6674 of LNCS, pages 73–86. Springer Berlin / Heidelberg, 2011.
[9] Mikhail Jiline, Stan Matwin, and Marcel Turcotte. Annotation concept synthesis and enrichment analysis:
a logic-based approach to interpretation of high-throughput experiments. In A. Farzindar and V. Keselj, editors,
Canadian Conference on Artificial Intelligence 2010, Lecture Notes in Artificial Intelligence 6085, pages 304–308,
Ottawa, May 31–June 2 2010.
[10] Étienne Ogoubi, David Pouliot, M. Turcotte, and Abdelhakim Hafid. Parallel multiprocessor approaches
to the RNA folding problem. In Parallel Processing and Applied Mathematics, pages 1230–1239, 2008.
[11] Étienne Ogoubi, Abdelhakim Hafid, and M. Turcotte. An Isometric on on-Chip Multiprocessor
Architecture. In Electronics, Circuits and Systems, 2007. ICECS 2007. 14th IEEE International Conference on,
pages 991–994, December 11–14, 2007 2007.
[12] M. Anwar and M. Turcotte. An approach to selecting putative RNA motifs using MDL principle.
In BIOCOMP’06 — The 2006 International Conference on Bioinformatics & Computational Biology, pages
560–565, Las Vegas, Nevada, USA, June 26–29 2006.
[13] M. Anwar and M. Turcotte. Evaluation of RNA secondary structure motifs using regression analysis.
In IEEE CCECE 2006 — Canadian Conference on Electrical and Computer Engineering, pages 1716–1721,
Ottawa, Canada, May 7–10 2006.
[14] T. Nguyen and M. Turcotte. Exploring the space of RNA secondary structure motifs using suffix arrays.
In S. Blair et al., editor, 6th International Symposium on Computational Biology and Genome Informatics
(CBGI 2005), pages 1291–1298, Salt Lake City, Utah, USA, July 21-26 2005.
[15] B. Masoumi and M. Turcotte. Simultaneous alignment and structure prediction of RNAs: Are three input
sequences better than two? In V.S. Sunderam, G.D. van Albada, P.M.A. Sloot, and J. Dongarra, editors, 2005
International Conference on Computational Science (ICCS 2005), Lecture Notes in Computer Science 3515,
pages 936–943, Atlanta, USA, May 22-25 2005.
[16] V. Jin and M. Turcotte. Detecting localized interspersed motifs in genomic sequences. In IMTC/05 IEEE
Instrumentation and Measurement Technology Conference, pages 267–270, Ottawa, Canada, May 17-19 2005.
[17] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. Application of inductive logic programming to discover
rules governing the three-dimensional topology of protein structure. In C. D. Page, editor, Proc. of the 8th
International Workshop on Inductive Logic Programming (ILP-98), Lecture Notes in Artificial Intelligence 1446,
pages 53–64, Berlin, 1998. Springer-Verlag.
Abstracts and/or papers read
[1] Aseel Awdeh, Marcel Turcotte, and Theodore J. Perkins. Cell type specific binding preferences of
transcription factors. In Great Lakes Bioinformatics Conference, GLBIO 2021, May 10–13 2021.
[2] Aseel Awdeh, Marcel Turcotte, and Theodore J. Perkins. WACS: Improving peak calling by optimally
weighting controls. In Great Lakes Bioinformatics Conference, GLBIO 2019, May 19–22 2019.
[3] Andrew Sowinski, Marcel Turcotte, Gilbert Arbez, and David Taylor. One-minute quizzes to identify
potential students atc risk in engineering courses. In Proceedings of the Canadian Engineering Education
Association (CEEA), University of Toronto, June 4–7 2017.
[4] Mikhail Jiline, Stan Matwin, and Marcel Turcotte. Annotation concept synthesis and enrichment analysis:
A logic-based approach to the interpretation of high-throughput biological experiments. In 2012 Learning
Workshop, Snowbird, Utah, April 3-6 2012. Computational and Biological Learning Society and the NIPS.
[5] S. Moreira, S. Breton, M. Valach, M. Aoulad Aissa, M Turcotte, and G Burger. Identification of an
elusive trans-splicing machinery. In Annual meeting of the Society for Molecular Biology & Evolution (SMBE
2012), Dublin, Ireland, June 23-26 2012. Poster.
[6] Sandrine Moreira, Marcel Turcotte, and Gertraud Burger. In silico identification of the elusive
trans-splicing machinery of a highly divergent protozoan. In 2011 MonBUG Bioinformatics Symposium,
Montréal, September 23 2011.
[7] Sandrine Moreira, Marcel Turcotte, and Gertraud Burger. Diversité géomique : le curieux mode
d’expression des gènes d’un eucaryote unicellulaire. In Association francophone pour le savoir (ACFAS),
Sherbrooke, Canada, May 9–13 2011.
[8] Georgette Kiethega, Marcel Turcotte, and Gertraud Burger. Nouveau mécanisme de trans-splicing dans la
mitochondrie des diplon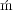 ides. In Association francophone pour le savoir (ACFAS), Sherbrooke, Canada, May
9–13 2011.
ides. In Association francophone pour le savoir (ACFAS), Sherbrooke, Canada, May
9–13 2011.
[9] Oksana Korol and Marcel Turcotte. Mining for relationships between biological markers in
high-throughput sequence data. In 2011 MonBUG Bioinformatics Symposium, Montréal, September 23 2011.
[10] Gertraud Burger, Georgette Kiethega, and Marcel Turcotte. Unconventional trans-splicing and RNA-editing
in mitochondria of an Euglenozoan. In ICPMB 2011. International Conference for Plant Mitochondrial Biology,
Hohenroda, Germany, May 14–19 2011.
[11] Oksana Korol and Marcel Turcotte. Module Inducer — a tool to automatically extract knowledge from
biological sequences. In RECOMB2011, 15th Annual International Conference of Research in Computational
Molecular Biology, Vancouver, March 28–31 2011.
[12] Yifei Yan, Seyed Amir Malekpour, Marcel Turcotte, Georgette Kiethega, and Gertraud Burger. RNA
editing and trans-splicing directed by gRNAs? In 2011 Gordon Research Conference on RNA Editing, January
9–14 2011. Poster.
[13] Sandrine Moreira, Marcel Turcotte, and Gertraud Burger. Analysis of potential nuclear RNA editing
sites in a highly divergent protozoa. In 2010 Robert Cedergren Bioinformatics Colloquium, 2010.
[14] Seyed Amir Malekpour, Marcel Turcotte, and Gertraud Burger. gRNA mediated trans-splicing of gene
modules in diplonema papillatum. In 2010 Robert Cedergren Bioinformatics Colloquium, 2010.
[15] Georgette Kiethega, Marcel Turcotte, and Gertraud Burger. cox1 gene fragmentation and rna editing in
diplonemid mitochondria. In 18th meeting of International Society for Evolutionary Protistrogy (ISEP XVIII),
Kanazawa, Japan, July 2–7 2010 2010.
[16] S. Kannan, M. Turcotte, and M. Burger. Automated (RNA) motifs discovery in the mitochondrial
genome of diplonema papillatum. Oral presentation, Robert Cedergren Bioinformatics Colloquium 2008,
Montréal, Canada, 2008.
[17] M. Jiline, K. Baetz, M. Turcotte, and S. Matwin. Knowledge enriched mining of systematic genome
screens using inductive logic programming. In Progress in Systems Biology 2006, Ottawa, November 9 and 10
2006. Oral presentation.
[18] S. Baird, M. Turcotte, R. Korneluk, and M. Holcik. Discovery of novel internal ribosome entry site motifs
from searches with the XIAP IRES structure. 2006 Cold Spring Harbor Laboratory Meeting on Translational
Control, Poster, September 6 – 10 2006.
[19] S. Baird, M. Turcotte, R. Korneluk, and Holcik M. Searching for IRES structurally similar to the XIAP
IRES. In RNA 2005, the Tenth Annual Meeting of the RNA Society, Banff, Canada. Poster #566, May 24-29
2005.
[20] S. Baird, M. Turcotte, R. Korneluk, and Holcik M. Searching for IRES. CIHR National Student Research
Poster Competition, Winnipeg, Canada, Poster Session, June 7-8 2005.
[21] B. Masoumi and M. Turcotte. A dynamic programming algorithm for [the] simultaneous alignment and
structure prediction of 3 RNA sequences. BioNorth 2004 — 11th Annual Ottawa Life Sciences International
Conference & Exhibition, Poster #114, Nov 29-Dec 1st 2004.
[22] S. Baird, M. Turcotte, R. Korneluk, and Holcik M. Structural determination of internal ribosome entry
sites with currently available software. In RNA 2004, the Ninth Annual Meeting of the RNA Society, Madison,
Wisconsin, Poster Session, June 1-6 2004.
[23] S. Baird, M. Turcotte, R. Korneluk, and M. Holcik. Searching for IRES. In The Canadian Genetic
Diseases Network Annual Scientific Meeting, Kimberly, Ontario, Poster Session, May 27-30 2004.
[24] M. Turcotte. Détermination de structures secondaires conservées dans les ARNs: Application à l’étude des
transcrits du gène c-myc des mammifères. 72e congrès de l’ACFAS, Montréal, Canada, May 10-14 2004. Oral
Presentation.
[25] S. Baird, S. Balabanian, R. G. Korneluk, M. Turcotte, and M. Holcik. Unique sequence characteristics
of internal ribosome entry sites useful for database search. BioNorth 2002 — 9th Annual Ottawa Life Sciences
International Conference & Exhibition, Poster session, November 4–6 2002.
[26] M. Turcotte. Automatic discovery of structural motifs. First Canadian Working Conference on
Computational Biology (CCCB’00), November 12 2000. Oral Presentation.
[27] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. Generating protein three-dimensional folds signatures
using inductive logic programming. In 2000 Convention of the Society for the Study of Artificial Intelligence
and the Simulation of Behaviour, Birmingham, UK, April 17-20 2000. Oral Presentation.
[28] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. Learning protein structure principles. In The 17th
Machine Intelligence Workshop, Suffolk, UK, July 19-21 2000. Oral Presentation.
[29] M. J. E. Sternberg, P. A. Bates, L. A. Kelley, R. M. MacCallum, A. Müller, S. Muggleton, and
M. Turcotte. Exploiting protein structure in the post-genome era. In Intelligent Systems for Molecular Biology
1999, 1999. Oral Presentation.
[30] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. Learning rules which relate local structure to specific
protein taxonomic classes. In The 16th Machine Intelligent Workshop, King’s Manor, York, December 1–3 1998.
[31] M. Turcotte, S.H. Muggleton, and M.J.E. Sternberg. A Prolog database unifying taxonomic and structural
information. In Intelligent Systems for Molecular Biology (ISMB 1998) Book of Abstracts, Poster Session,
Montréal, Canada, June 28–July 1 1998.
[32] M. Turcotte and M. Feeley. A parallel functional program for searching a discrete space of nucleic acid 3d
structures. In Dagstuhl-Seminar, Germany, 1994. Oral Presentation.
[33] F. Major, M. Turcotte, and G. Lapalme. Constraint satisfaction in functional programming. In Principles
and Practice of Constraint Programming, pages 174–177, Newport, Rhode Island, April 28–30 1993.
[34] M. Turcotte, G. Lapalme, and R. Cedergren. A measure of association for coordinated substitutions in
proteins. In Book of Abstracts, Patterns of Biological Organizations, Rensselaerville, NY, October 1992. 1992
Albany Conference. Poster Session.
[35] D. Gautheret, M. Turcotte, R. Cedergren, F. Major, and G. Lapalme. Modeling and prediction of RNA
3-D structures by combining symbolic and numerical approaches. J. Biomol. Struct. Dynam., 8(6):a061, June
1991.
[36] R. Cedergren, F. Major, D. Gautheret, E. Fillion, L. Jolicoeur, M. Turcotte, and R. Cedergren. A
3-D model of the hammerhead domain derived from experimental and topological constrainsts. International
Conference on Catalytic RNA as Anti-HIV agent: Design and Delivery to Cells, San Diego, California, October
1990. Oral Presentation.
[37] M. Turcotte, F. Major, G. Lapalme, D. Gautheret, E. Fillion, and R. Cedergren. Using functional and
logic programming in molecular graphics applications. 9th annual meeting of the Molecular Graphics Society,
helded Uppsala, Sweeden, Poster Session, July 1990.
[38] M. Turcotte, G. Lapalme, R. Cedergren, and F. Major. A user-interface for the folding and unfolding
system (FUS). 1st C.I.A.R. Evolution Group Meeing for Graduate Students, Université de Montréal, Montréal,
Canada, Oral Presentation, 1989.
Contributions to publications via supervisions
[1] Kevin Sutanto. RNA sequence classification using secondary structure fingerprints, sequence-based
features, and deep learning. Master of computer science, University of Ottawa, School of Electrical Engineering
and Computer Science, 2021.
[2] Manuel Belmadani. MotifGP: DNA motif discovery using multiobjective evolution. Master of computer
science, University of Ottawa, School of Electrical Engineering and Computer Science, 2016.
[3] Aseel Awdeh. The potential power of dynamics in epistasis analysis. Master of computer science, University
of Ottawa, School of Electrical Engineering and Computer Science, 2015.
[4] Alexander Gawronski. RiboFSM: Frequent subgraph mining for the discovery of RNA structures and
interactions. Master’s thesis, School of Electrical Engineering and Computer Science, University of Ottawa,
2013.
[5] Oksana Korol. ModuleInducer: automating the extraction of knowledge from biological sequences. Master’s
thesis, University of Ottawa, School of Electrical Engineering and Computer Science, Defended on September
20, 2011 2011.
[6] Mikhail Jiline. Annotation Concept Synthesis and Enrichment Analysis: a Logic-Based Approach to the
Interpretation of High-Throughput Biological Experiments. PhD thesis, Ottawa-Carleton Institute for Computer
Science, University of Ottawa, 2011.
[7] Predrag Mizdrak. Novel iterative approach to joint sequence alignment and tree inference under maximum
likelihood: a critical assessment. Master’s thesis, Ottawa-Carleton Institute for Computer Science, University of
Ottawa, 2008.
[8] Stephen Baird. Searching for IRES. PhD thesis, Human and Molecular Genetics Program, Biochemistry,
Microbiology and Immunology Department, Faculty of Medicine, University of Ottawa, 2006.
[9] Mohammad Anwar. Implementation and evaluation of scoring schemes for the automated discovery of
nucleic acid structures. Master’s thesis, Ottawa-Carleton Institute for Computer Science, 2006.
[10] Mohak Shah. Sample Compression, Margins and Generalization: Extensions to the Set Covering Machine.
PhD thesis, Ottawa-Carleton Institute for Computer Science, University of Ottawa, 2006.
[11] Beeta Masoumi. A dynamic programming algorithm for the simultaneous alignment and structure
prediction of three RNA sequences. Master’s thesis, School of Information Technology and Engineering, Faculty
of Engineering, University of Ottawa, 2005.
[12] Truong Nguyen. Efficient determination of concensus secondary structures in RNA. Master of applied
science in engineering, School of Information Technology and Engineering, Faculty of Engineering, University of
Ottawa, 2004.
[13] Victor Jin. A computational approach to the analysis of localized interspersed motifs in complete genomic
sequences. Master’s thesis, Ottawa-Carleton Institute for Computer Science, 2003.
Papers in preparation for referred journals
[1] Aminur Rab Ratul, Marcel Turcotte, M. Hamed Mozaffari, and WonSook Lee. Prediction of 8-state protein
secondary structures by 1D-Inception and BD-LSTM. bioRxiv, 2019.
Invited and public talks
-
Learning relationships between over-represented motifs in a set of DNA sequences. The First Bioinformatics
Scientific Meeting in KSU (BioSM-KSU). Invited Speaker. King Saud University (KSU), May 4, 2014.
-
Consensus RNA secondary structures determination. Université du Québec en Outaouais. February 24, 2010.
-
Automated (RNA) motifs discovery in the mitochondrial genome of Diplonema papillatum, University of
Ottawa, Faculty of Medicine, RNAClub. May 6, 2009.
-
Predicting the consensus RNA secondary structure for k > 2 sequences, University of Memphis, February 27,
2009.
-
Applying Relational Learning to Structural Molecular Biology Problems, IEEE Computational Intelligence
Society (CIS) — Ottawa Chapter, 2008/07/03.
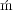 ides. In Association francophone pour le savoir (ACFAS), Sherbrooke, Canada, May
9–13 2011.
ides. In Association francophone pour le savoir (ACFAS), Sherbrooke, Canada, May
9–13 2011.